The history of the pea gene map: last revolutions and the new symbiotic genes
| Rozov, S.M., Kosterin, O., | Institute of Cytology and Genetics Novosibirsk 630090 Russia |
| Borisov, A.Y. and Tsyganov, V. | All-Russia Research Institute for Agric.
Microbiology |
The garden pea, Pisum sativum, was the model system used by Gregor Mendel to formulate his famous Mendel’s Laws. But the characters used by Mendel have been since shown to be controlled by genes belonging to different linkage groups or to distant parts of the same linkage group. Thus, Mendel failed to observe the phenomenon of linkage. Instead, linkage was first described and interpreted by Thomas Morgan in his works with Drosophila.
Linkage in pea was first described in 1912 by Vilmorin and Bateson (6). Thus, if the history of pea genetics begins with the Gregor Mendel, the history of the pea linkage map begins much later. Initially, only pairs and triplets of linked genes were described. As the amount of known genes increased, the first linkage groups were established. The first pea linkage map was constructed by Wellensiek in 1925 (7), who had established six linkage groups. Interestingly, in this first pea gene map gp belonged simultaneously to two independent linkage groups. Next, in 1948, Lamprecht (3) published his well-known full pea gene map with seven linkage groups. It was a glorious time for classical genetics and the chromosome theory of inheritance. In this paradoxical time any investigator – and Lamprecht of course – tried to produce a linkage map with the same number of linkage groups as there were chromosomes. Lamprecht’s map consisted of seven linkage groups, where a (anthocyanin) was on the end of the first chromosome. The fifth group contained gp, which on Wellensiek’s map was linked not only to te and cri, but also to r and tl of Lamprecht’s chromosome 7.
Lamprecht’s map was stable until the mid and late 1980’s when isozyme and morphological markers were placed on linkage groups that did not correspond any of Lamprecht’s groups (5, 8, 9). At the same time a large amount of chromosome gene material - equivalent to about 50 map units - was found above the gene a in the first linkage group (8, 9). Ian Murfet (4) described the gigas gene, and showed linkage to both to gp and tl - reestablishing Wellensiek’s linkage group. The new linkage group V was established as a result of a fusion of Lamprecht’s 5 and 7. This time was the beginning of a renaissance in the pea genetics – the global pea map rearrangement. The new linkage group VII was constructed with biochemical and morphological genes. Furthermore, our investigations indicated that the a and d segments of Lamprecht’s 1st chromosome were not linked (1, 2), and this group was divided into two conventional parts.
The next step in the pea linkage map revolution was made by the efforts of Norman Weeden, Noel Ellis and Ian Murfet. These researchers, using a great number of RFLP and RAPD markers, RIL-lines and some morphological, biochemical and physiological markers, succeeded in combining linkage groups IA and II and demonstrating that the Np-le segment was on linkage group III, not on IV. Linkage group IB now appears to represent chromosome 1.
In spite of these brilliant mapping efforts, there remain some ambiguities in the existing pea linkage map. In some crosses we have observed a reliable linkage between genes on linkage group IA with some markers on linkage group VI. It is not impossible that in the next years linkage group IA will be fused with the linkage group VI rather than II. Moreover, some evidence exists that there is a large amount of chromosome material out of the boundaries of the linkage group IB. Thus, in our opinion, the pea gene map revolution is not finished, and the next few years will be no less fruitful then the previous.
As this workshop is dedicated mainly to the pea symbiosis genes, some mention about the position of the sym genes on the pea map must be made. Most of the sym genes were localized during the last twenty years, but their relative positions on the map have been quite stable. Fifteen years ago, two loose clusters of sym genes in linkage group I and randomly dispersed sym genes on the other chromosomes were reported. The Fabaceae family is believed to have originated at the beginning of the Cretaceous. Even if these genes were clustered at the eve of the rhizobium symbiosis formation, 70 millions years is quite enough to disperse them throughout the genome. Moreover, the rhizobium symbiosis gene machinery is now believed to have arisen from more ancient mycosymbiosis gene systems, which evolutionary age is more then 400 millions years. We believe that the clustering of sym linkage group I is only an accident. What sort of a functional cluster could it be if the genes are separated by 5 to 10 percent recombination?
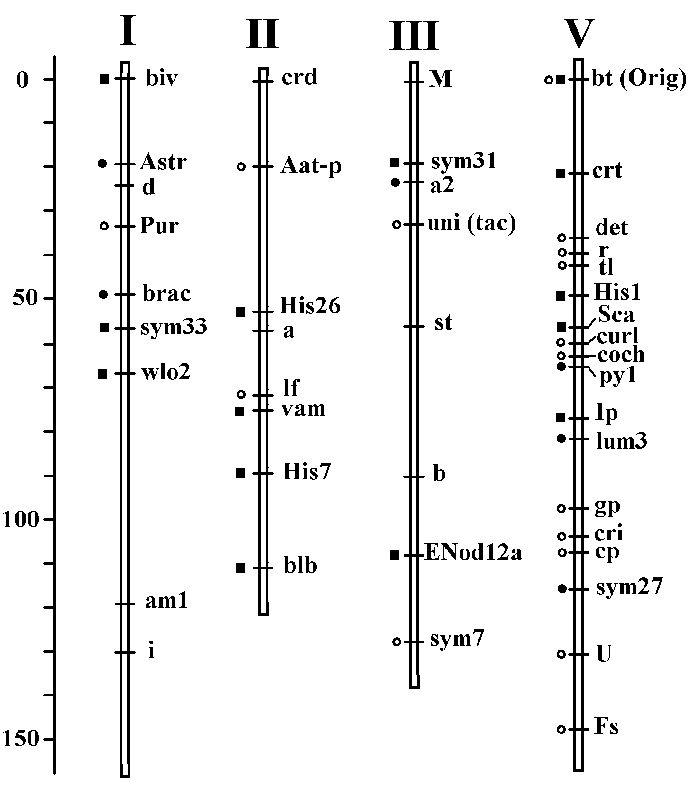
During the last 15 years our genetic, biochemical and evolutionary studies of Pisum have given us the new material for modification and further development of the existing pea linkage map. During this period we have described and localized 13 new genes, and 6 previously identified genes were put on the map. The positions of a large amount of previously localized genes were improved and verified in our studies. The main results of this work is illustrated on the next figure:
The new genes, found and localized by us are marked on this picture with filled squares, genes initially localized by us - with filled circles, and the genes, with positions confirmed in our work - with the empty circles.
We work mainly with the genes of four linkage groups, now referred to as I, II, III and V. A large number of new morphological markers - biv, wlo2, vam, blb, crt, were mapped, some biochemical markers, among which three histone H1 loci, a protease inhibitors gene cluster and seed albumin Sca, and four symbiosis genes: sym27, sym31, sym33 and Enod12.
The data seen on this picture represent a new version of corresponding chromosome map segments, and now it is integrated in the last official version of the pea linkage map.
 |