Genetic instability in the histone H1 locus of pea (Pisum sativum L.)
Bogdanova, V.S., Trusov, Y.A. Institute
of Cytology and Genetics
and Berdnikov, V.A. Novosibirsk
630090, Russia
Instability of expression of genetic loci is usually represented by variegated phenotypes as sectors of wild-type tissue on the otherwise mutant background or vice versa. The phenomenon is rather frequent in plants and can be observed in the sectored and spotted patterns of flower or chlorophyll pigmentation. An unstable behaviour of mutable alleles with high frequency of reversion is usually attributed to the insertion and excision of transposable elements (17). Many unstable alleles of different loci in maize and some alleles in snapdragon were molecularly and genetically characterized and shown to harbour controlling elements (reviewed in 8).
Transposon-like sequences were described in other plant species such as Glycine max (23), Triticum aestivum (11), Petroselinum crispum (13), and Pisum sativum (4, 21). However, these sequences were not found to transpose or to be connected with genetic instability. On the other hand, there is a large number of mutable loci described in various plant species which are, as implied by genetic analysis, associated with transposable elements (see for example, 5 and references therein). Lack of appropriate molecular probes hinders establishing of an unambiguous connection between genetic instability and transposition of a certain element.
Histone H1 of pea is encoded by at least three genetic loci (16). The gene of the most abundant subtype 1 (His1) is localized in linkage group V (20), the gene of subtype 7 (His7) is in linkage group IA (15), and the genes of subtypes 2 to 6 (His2-His6) are tightly linked (2) and also located in group IA as a gene cluster named His (2-6) (1, 16). In the present paper we describe a phenomenon of unstable expression of a gene encoding one subtype of histone H1 in pea, the mutable allele being isolated from a collection of germplasm of the All-Russian Plant Breeding Institute (VIR).
Materials and methods
Plant material was originally obtained from the All-Russian Plant Breeding Institute, St. Petersburg, Russia and from the Nordic Gene Bank, Alnarp, Sweden. This material was used in our laboratory to construct a series of early-ripening lines, which were included in our analysis.
Designation of allelic combinations, or haplotypes, of H1 subtypes 3-6 was as accepted in (2) (the allelic state of subtype 2, belonging to the same cluster, was not included because its “normal” allele predominates overwhelmingly). The haplotype was designated by a formula consisting of a combination of four digits. The first position in the formula refers to subtype 3, the second to subtype 4 and so on. The digit itself reflected the allelic state of the relevant subtype. Alleles of each gene were numbered according to the increase of electrophoretic mobility of the encoded protein, 1 referring to the slowest variant, 2 to the second slowest and so on. 0 indicates the absence of a corresponding electrophoretic band.
Histone H1 was isolated by Johns’ (14) method with further modifications. About 200-500 mg of pea leaves were homogenized in 10 ml of 0.15 M NaCl, the homogenate was filtered through a grid (1 ґ 1 mm2) of stainless steel and centrifuged at 1500 g for 5 min. Histone H1 was extracted by resuspending the pellet in 1 ml of 5% HClO4. After centrifugation, the protein was recovered from the supernatant by adding 6 volumes of cold acetone and sulphuric acid to a concentration of 0.5 M. The precipitated protein was centrifuged and then dissolved in 0.02 ml of a medium containing 0.9 M acetic acid, 8 M urea, and 15% sucrose. The preparations were subjected to electrophoresis in long (up to 40 cm) slabs of 15% polyacrylamide/0.5% N,N’-methylenbisacrilamide gel containing 8 M urea and 0.9 M acetic acid following a modification (3) of Panyim and Chalkley’s (19) method. After electrophoresis, the gels were stained in 0.01% Coomassi R-250 in 0.9 M acetic acid.

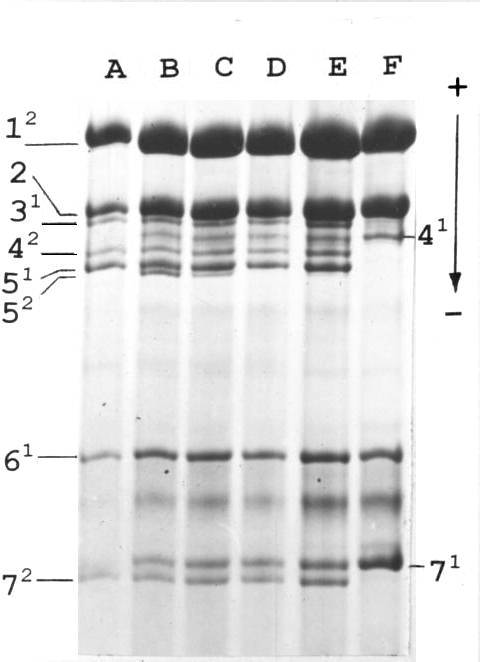
Fig. 1. Histone H1 of parental lines and F1 hybrids. A: tester line (haplotype 1211); B – E: F1 hybrids; F – VIR-4362 (haplotype 0101). Numerals stand for subtypes of histone H1, superscripts for their allelic variants.
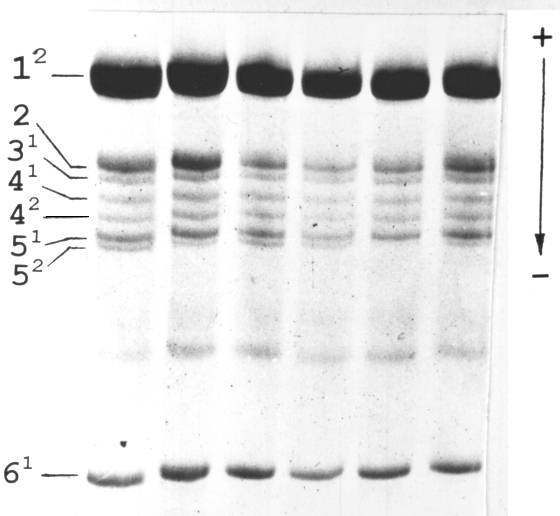
Fig. 2. Histone H1 isolated from leaves from different nodes of a single pea plant.
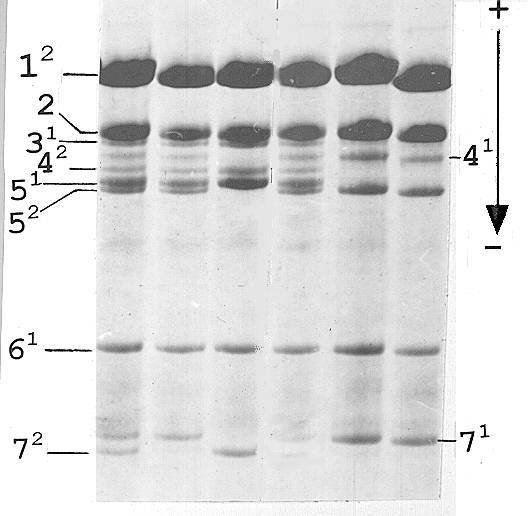
Fig. 3. Histone H1 of the F2 family derived from the F1 plant with histone H1 shown in Fig. 1, lane B.
Results
In our previous work (2) we have performed an extensive analysis of histone H1 of peas originating from different regions of the Old World. Among 883 analysed accessions we found one, VIR-4362, originating from the Vologda region that possessed an unusual haplotype with an absence of subtypes 3 and 5 (haplotype 0101). One plant with the haplotype 0101 was crossed (as a female parent) to a tester line from our collection possessing haplotype 1211 (in our designation “Sprint-1211”). Unexpectedly, in the F1 generation we did not observe a uniformity of histone H1 phenotype. Instead of plants heterozygous for subtype 4 (second position in the formula) and hemizygous for subtypes 3 and 5 (looking like homozygote for allele 1 of subtype 3 and homozygote for allele 1 of subtype 5) we found F1 hybrids displaying two electrophoretic bands corresponding to subtype 5, namely to its allelic variants 1 and 2 (Fig. 1), the latter being absent from both parents. The second unusual feature of these hybrids was the variable level of expression of the novel allele 2. It varies from zero (Fig. 1, lane D) through weak (Fig. 1, lanes E and C) up to the level of expression of a tester allele 1 (Fig. 1, lane B) forming a common heterozygous phenotype. We named the allele of subtype 5 coming from the VIR-4362 stock the “Vologda” allele.
We isolated histone H1 from different parts of the same F1 plant to analyse the distribution of activity of the Vologda allele. As shown in the Fig. 2 we observed intraplant variability of its expression.
Segregation of alleles of subtype 5 in the F2 generation was as follows. F1 plants with equal expression of the Vologda allele and tester allele 1 (Fig. 1, lane B) gave rise to a progeny where all chromosomes carrying the His2-6 locus from the VIR-4362 parent were of the 0121 haplotype both in homozygotes and in heterozygotes (Fig 3), thus indicating that an event of activation of the Vologda allele took place in the germ line tissue and generated a stable active derivative. All the other F1 plants resulted in progeny with variable expression of the Vologda allele resembling that of the F1 generation.
We analysed expression of allele 2 of subtype 5 in the VIR-4362 stock. Among 328 plants only one was found to display a high level of expression while others either lacked the corresponding band, or it was hardly detectable and distinguishable only in heavily loaded samples. Thus, occurrence of the high level of expression of the Vologda allele was met much more rarely than after cross-pollination – one case among 328 plants analysed compared with 2 plants of 10 tested in F1.
It is interesting to note that the only plant with high expression of the Vologda allele appeared to be a heterozygote since in its progeny we observed 0101 haplotypes, intermediates and 0121 haplotypes. The proportion of 12 plants of haplotype 0101 out of 39 tested corresponds well to 1/4 (Chi-sq. 0.69, P > 0.7).
Discussion
Our wide experience, in analysing the histone H1 spectrum in peas (16) leads us to conclude that a novel band appearing in F1 plants, which is absent in both parental lines, cannot be a post-translational modification of any histone H1 subtype. In very young plant organs, we observed only some modification of some subtypes, most probably phosphorylation, yielding a series of a few diffuse bands of decreasing intensity with slowed electrophoretic mobility (16). On the other hand, the new band corresponds exactly to the well known allelic variant 2 of H1 subtype 5.
The simplest implication for the observed activation of the “latent” allele after crossing is to suggest that expression of the Vologda allele in the haplotype 0101 is “turned off” by an insertion of a controlling element. After cross-pollination, a transposon activates and begins to excise with resulting restoration of expression of the Vologda allele. In the light of this suggestion it is reasonable to expect that heterozygous plants displaying weak expression of allele 2 are the mixtures of cell clones with active and inactive Vologda allele. As shown in the Fig. 2, different parts of the plant vary in the intensity of the electrophoretic band corresponding to the Vologda allele. Thus, we can speak of a variegated phenotype in respect of expression of the subtype 5 gene His5 of histone H1.
The mode of expression of the Vologda allele in the VIR-4362 stock is in good accordance with the assumption of a transposable element residing in the His5 gene. Weak expression appears to be a consequence of a somatic reversion while absence of detectable expression may be due to the fluctuation of the activating factor(s)’ concentration and its decrease below some threshold level.
We observed some cases of both somatic and germinal reversion to a high level of expression of the His5 gene. Germinal reversion generated stably expressed derivative corresponding to the 2 allele of subtype 5 detectable just after cross-pollination in the F1 and in the subsequent generation. Unfortunately, due to the insufficient number of F1 plants available, we cannot estimate the frequency of germinal reversion. However, we can anticipate it to be rather high (2 plants of 10 tested). Preliminary data suggest that stable null derivatives also occur.
Interestingly, in 17 F1 hybrids from the crosses of the VIR-4362 plant as a pollen parent with tester lines of 1133 and 1211 haplotypes, we did not observe detectable expression of the Vologda allele. However, in F2 progeny from these crosses activation of the Vologda allele was apparent. Dependence of the maize Spm element transposition frequency on the direction of the cross was observed in some cases (7, 18).
The nature of activating factors responsible for the transpositions of the element remains unclear. It may be another transposon – an active member of a two-component system (reviewed in 10) or some chromosomal gene coding for transposase. We know a number of genes in pea which condition variegated patterns of a seed coat coloration such as F and Fs (violet spots), M (marbled patches), and Ust (purple stripes) (6). There are unstable alleles affecting other parts of the plant, for example, And produces purple spots on leaves (22) and Pur (purple pod) is known to mutate frequently to weaker alleles (6). The products encoded by these genes are not known, however, one can speculate that these genes code for a kind of transposase which could be utilized by the transposon residing in the His5 gene. At least, they could serve as markers responding to the presence of transposase, of course, if these genes and the His5 gene contain related transposons. It should be noted that the VIR-4362 stock lacks detectable coloration of the seed coat while the majority of peas from the VIR collection display either F/Fs spotting or M marbling, or both.
Inversely, VIR-4362 may contain a gene responsible for a repressor of transposition like the st gene of Antirrhinum majus (12). This is the subject of further genetic analysis.
In conclusion it should be noted that the presence of a putative transposon in the His5 gene provides a good opportunity for isolation and molecular characterization of the element since the DNA sequence of a histone H1 gene from pea is determined (9). In turn, this opens the way for transposon tagging and further investigation of the pea genome.
Acknowledgement. This work was partly supported by the Russian National Program “Basic Research Foundation”.
* * * * *