|
Pisum Genetics |
Volume 24 |
1992 |
Research Reports |
pages 32-36 |
The genetic basis of early flowering in mutant line I/178
|
Arumingtyas, E.L. and Murfet, I.C.
|
Department of Plant Science University of Tasmania Hobart, Tasmania 7001, Australia |
We report here on the genetic identity of the early flowering mutant, I/178, obtained by Dr M. Vassileva (Genetics Institute, Sofia) following treatment of cv Ramonsky 77 with 100 r Nf. Seeds of I/178 and Ramonsky 77 were provided by Dr Stig Blixt under the Weibullsholm catalogue numbers WL6028 and WL2164, respectively.
The tests were conducted in the phytotron at Hobart. The phenotypes of the mutant and its initial line were determined by comparison with standard lines from the Hobart collection representing several flowering classes defined previously (3, 8). The long day photoperiods used were 18 h (natural daylight extended before dawn and after dusk by fluorescent and incandescent light providing 25 mmol m-2 s-1 at pot top) or 24 h (8 h daylight + 16 h incandescent light providing 3 mmol m-2 s-1 at pot top). The short day photoperiod used was 8 h (8 h daylight + 16 h dark). Lateral shoots were excised regularly and results were collected only from primary shoots. Plants represented in Figs 1 and 3 were transferred from an 8 h to an 18 h photoperiod at age 82 and 56 days, respectively. All plants had produced either open flowers or macroscopically visible flower buds at the time of transfer. The node of flower initiation was taken as the first node to bear a flower initial regardless of whether or not the initial developed into an open flower. Node counts commenced from the first scale leaf as 1. Flowering time was taken as the number of days from sowing to first open flower. Further details on growing conditions and scoring procedures are given in previous papers (8, 10, 11). Details of the major flowering genes in pea may be found in a recent review (9).
Our tests showed the initial line Ramonsky 77 (WL2164) has a similar flowering phenotype to Hobart line 24 (Figs 1-3), the standard line for the L flowering class, i.e. Ramonsky is late flowering with a quantitative response to photoperiod. Ramonsky generally commenced flowering at nodes 14-16 under long day conditions (18 or 24 h) and in the range of nodes 23-30 under short day conditions (8 h). Mutant I/178 has a similar flowering phenotype to Hobart line 60 (Figs 1-4), the standard line for the EI flowering class. Typical of plants in the EI class, line I/178 commenced flower initiation in the range of nodes 9-13 regardless of photoperiod but days to first open flower and number of reproductive nodes were significantly increased under short day conditions.
Given what is known about the action and interaction of the major flowering genes in pea (see 9) it seems likely from the phenotypic data that Ramonsky 77 has a flowering genotype of Lf E Sn Dne hr and that mutant I/178 has resulted from a mutation of the type Lf to lf. A less likely possibility is that Ramonsky 77 has genotype lf e Sn Dne hr and the mutation was of the type e to E. The results for the F1 and F2 of cross I/178 x Ramonsky 77 (Fig. 2) support the first alternative since they indicate monogenic recessive inheritance of the mutant trait. The observed numbers of 22 late and 9 early plants are in good accordance with a 3:1 ratio (c2 = 0.27).

Fig. 1. Node of flower initiation and flowering time for WL2164 (Ramonsky 77), mutant I/178 and standard lines L59 (lf E sn Dne hr), L73 (Lf E sn Dne hr), L60 (lf E Sn Dne hr) and L24 (Lf e Sn Dne hr) representing the flowering classes ED, DN, EI and L, respectively. Photoperiod 8 h.
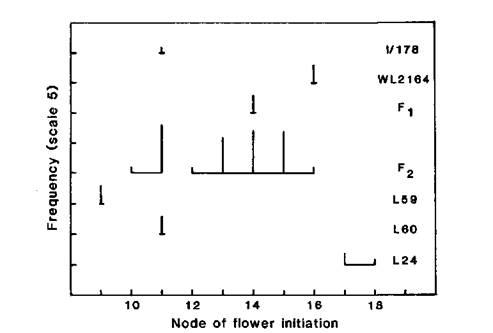
Fig. 2. Frequency distribution for node of flower initiation for lines WL2164 (Ramonsky 77), mutant I/178, the F1 and F2 of cross WL2164 x I/178, and standard lines 59, 60 and 24. Photoperiod 18 h.
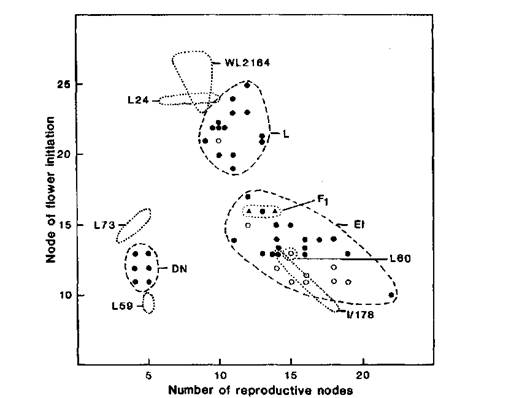
Fig. 3. Node of flower initiation and number of reproductive nodes for WL2164 (Ramonsky 77), mutant I/178, standard lines 59 (phenotype ED, lf E sn Dne hr), 73 (DN), 60 (EI, lf E Sn Dne hr) and 24 (L, Lf e Sn Dne hr), and the F1 (▲) and F2 of cross I/178 (proposed genotype a lf E Sn Dne hr) x L73 (A Lf E sn Dne hr). Red (A-) and white (aa) flowered F2 segregates are identified by closed (●) and open (○) circles, respectively. Photoperiod 8 h.
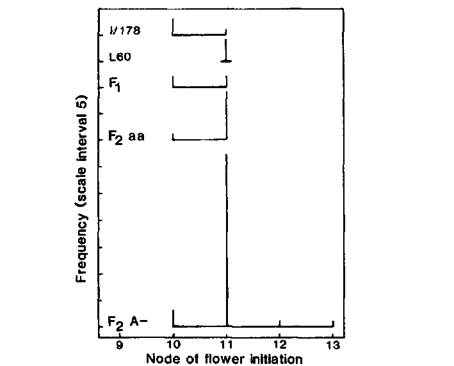
Fig. 4. Node of flower initiation for lines I/178 and 60, and the Fj and F2 cross I/178 x L60. F2 data are plotted separately for red (A-) and white (aa) flowered plants. The parents and F1 were planted March 11 and the F2 on May 30. Photoperiod 8 h.
Two further genetic tests were conducted by crossing mutant I/178 with lines of known genotype. Both tests utilised the fact that Lf is located on chromosome 1 about 10 cM from gene A, a basic gene for anthocyanin production (1, 2, 4, 5, 15). Thus A serves as a fairly effective marker to follow segregation at the Lf locus. The mutant and its initial line have white (a) flowers.
Data for the F2 of cross I/178 (proposed genotype a lf E Sn Dne hr) x L73 (A Lf E sn Dne hr) are shown in Fig. 3. Line 73 has a DN (day neutral) flowering phenotype since it carries the recessive sn allele. It is apparent that the majority of the white flowered F2 plants had a flowering phenotype similar to line I/178. This is consistent with the proposed genotype for I/178 and the coupling phase arrangement for a and lf. Lf is epistatic to E (4) and although neither parent in this cross flowered later than node 16, some late segregates with a phenotype like Ramonsky 77 occurred in the F2 as a result of this interaction and restoration of the proposed initial genotype Lf E Sn Dne hr. Indeed if Lf were fully dominant the F1 should have phenotype L like Ramonsky and the F2 should segregate in the ratio of 9L:3EI:4DN. However, experience has shown (4, 10) that it is not uncommon for heterozygous plants of genotype Lf lf EE Sn- DneDne hrhr to flower within the EI rather than the L class. Such was the case here where the coordinates for the two Fl plants plot within the EI class near the upper bound (Fig. 3). Again, the observed F2 numbers of 15 L, 26 EI and 6 DN plants deviate significantly (P < 0.001) from those expected on the basis of a 9:3:4 ratio, largely as a result of a deficiency of L-type and an excess of EI-type segregates. The situation can be explained if most of the red-flowered EI segregates have genotype Lf lf and this explanation is consistent with the fact that the majority of red flowered EI segregates flower at a higher node than both line I/178 and the majority of white flowered EI segregates. The observed number of DN segregates in the F2 is also less than that expected but the deficiency is not statistically significant (P > 0.05) and Sn-sn segregations are frequently closer to a 4:1 than a 3:1 ratio (4).
As expected, the cross between the two EI lines I/178 (a lf E Sn Dne hr) and 60 (A lf E Sn Dne hr) produced only EI offspring in F1 and F2; the parents, Fl and F2 all flowered within the narrow range of nodes 10-13 (Fig. 4). The red (A-) and white (aa) flowered F2 segregates commenced flower initiation at nodes 11.00 ± 0.08 (n=37) and 10.91 ± 0.09 (n=11), respectively. These two means are not significantly different (P > 0.5).
We conclude that Ramonsky 77 has genotype Lf E Sn Dne hr and that mutant I/178 is the result of mutation of Lf to an allele of very similar strength to the lf allele present in line 60, the standard EI line.
Four naturally occurring Lf alleles have been identified - Lfd, Lf, Lf and lfa (4, 5). The present case brings to 21 the number of induced mutations traced to the Lf locus (6, 7, 10-14). Thus the Lf gene appears to be some 10-20 times more susceptible to mutation than the other major flowering genes.
Acknowledgements. We thank Peter Bobbi and Katherine McPherson for technical assistance and the Australian Research Council and the International Development Programme for financial support.
Hoshino, Y. 1915. J. Coll. Agric. Hokkaido Imp. Univ. 6:229-288.
Lamprecht, H. 1961. Agri Hort. Gen. 19:360-401.
Murfet, I.C. 1971. Heredity 26:243-257.
Murfet, I.C. 1971. Heredity 27:93-110.
Murfet, I.C. 1975. Heredity 35:85-98.
Murfet, I.C. 1978. PNL 10:48-52.
Murfet, I.C. 1982. In Documentation of Genetic Resources : a Model, Eds S. Blixt and J.T. Williams, IBPGR, Rome, pp 45-51.
Murfet, I.C. 1985. In Handbook of Flowering, Vol. IV, Ed A.H. Halevy, CRC Press, Boca Raton, Florida, pp 97-126.
Murfet, I.C. 1990. PNL 22:78-86.
Murfet, I.C. 1991. Pisum Genetics 23:16-18.
Murfet, I.C. and Ezhova, T.A. 1991. Pisum Genetics 23:19-25.
Murfet, I.C. and Groom, K. 1984. PNL 16:57-58.
Uzhintseva, L.P. and Sidorova, K.K. 1979. Genetika 15:1076-1082.
Uzhintseva, L.P. and Sidorova, K.K. 1988. PNL 20:39-40.
Weeden, N.F., Ambrose, M. and
Swiecicki, W.K. 1991. Pisum Genetics 23:
cover.